Sensitivity Analysis
Sensitivity analysis is the study of how the variations in the outputs of a system can be assigned to different sources of variation in its inputs.
Understanding the variance of the outputs of a system in terms of the contributions of the inputs is a key aspect of uncertainty quantification: it helps to test the robustness of a system, design strategies for uncertainty reduction and helps to calibrate physical models to the real world by improving their interpretability.
Sensitivity analysis methods are divided into two main classes: local methods, where changes in the output are studied for specific values of the inputs, and global methods where the total variability of the outputs is assigned to each input variable. Given a physical model of a system, for instance a climate model simulator, local sensitivity analysis is carried out using gradients. However, if the physical model is expensive-to-evaluate approximating gradients becomes intractable. Using an emulator of the physical model, like a Gaussian process, and using its analytic gradients is an effective alternative.
A similar situation arises when doing global sensitivity analysis. The core element in global sensitivity analysis are the Sobol indices, also called main effects [1,2]. They are a measure of “first order sensitivity” of each input variable. They account for the proportion of variance of the output explained by changing each variable alone while marginalizing over the rest. The total effects are also common [3]. They account for the contribution to the output variance of each variable but including all variance caused by the variable alone and all its interactions of any order of the inputs. Both indices can be easily computed in Emukit.
To illustrate how to do global sensitivity analysis of a simulator we use the Ishigami function. This function has three continuous inputs and depends on two parameters that we fix in this example.
from emukit.test_functions.sensitivity import Ishigami
ishigami = Ishigami(a=5, b=0.1)
target_simulator = ishigami.fidelity1
Emukit allows to compute both the main and total effects using Monte Carlo with the Saltelli estimators [4]. Instead of computing the indices by calling the simulator, we learn an emulator first and we compute the indices using cheap calls to this model. We first define the parameter space:
import numpy as np
from emukit.core import ContinuousParameter, ParameterSpace
space = ParameterSpace([ContinuousParameter('x1', -np.pi, np.pi),
ContinuousParameter('x2', -np.pi, np.pi),
ContinuousParameter('x3', -np.pi, np.pi)])
Now we generate a data set by evaluating the simulator on 500 random samples in the input domain:
from emukit.experimental_design.model_free.random_design import RandomDesign
desing = RandomDesign(space)
X = desing.get_samples(500)
Y = ishigami.target_simulator(X)[:,None]
And we wrap and fit a Gaussian process to the inputs and outputs of the simulator.
from GPy.models import GPRegression
from emukit.model_wrappers import GPyModelWrapper
model_gpy = GPRegression(X,Y)
model_emukit = GPyModelWrapper(model_gpy)
model_emukit.optimize()
We are now ready to compute the sensitivity indices. It is as simple as running:
from emukit.sensitivity.monte_carlo import MonteCarloSensitivity
senstivity = MonteCarloSensitivity(model = model_emukit, input_domain = space)
main_effects, total_effects, _ = senstivity.compute_effects(num_monte_carlo_points = 10000)
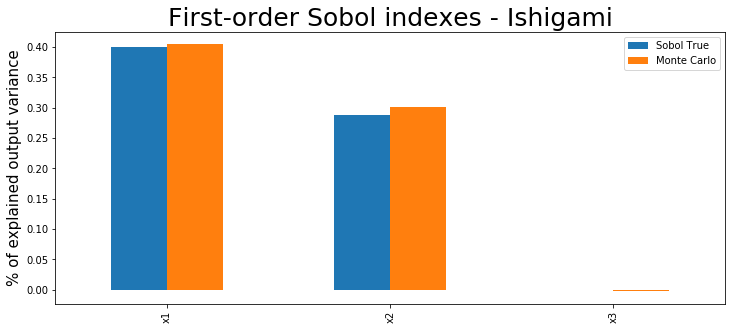
And that’s it! You can check the computed main and total effects. Note that although we have used a Gaussian process here, the coefficients are computed running Monte Carlo, which means that any model can be wrapped here.
Check our list of notebooks and examples if you want to learn more about how to do sensitivity analysis and other methods with Emukit. You can also check the Emukit documentation.
We’re always open to contributions! Please read our contribution guidelines for more information. We are particularly interested in contributions regarding examples and tutorials.
References on sensitivity analysis
- [1] Sobol, I. M. (2001), Global sensitivity indices for nonlinear mathematical models and their Monte Carlo estimates. Math Comput. Simulat., 55(1–3),271-280, 2001
- [2] Sobol’, I. M (1990). Sensitivity estimates for nonlinear mathematical models. Matematicheskoe Modelirovanie 2, 112–118. in Russian, translated in English in Sobol’ , I., 2001.
- [3] Andrea Saltelli, Paola Annoni, Ivano Azzini, Francesca Campolongo, Marco Ratto, and Stefano Tarantola. (2010) Variance based sensitivity analysis of model output. Design and estimator for the total sensitivity index. Computer Physics Communications, 181(2):259-270, 2010.
- [4] Kennedy, M.C. and O’Hagan, A., (2000). Predicting the output from a complex computer code when fast approximations are available. Biometrika, 87(1), pp.1-13, 2000.