Experimental design
Experimental design addresses the problem of how to collect data points (experiments) to better control certain sources of variance of a model.
In experimental design the goal is to decide at which locations of the input space we should evaluate a function of interest. In some contexts it is also known as active learning, for instance in image classification problems in which more labels need to be collected.
The are two main ways of doing experimental design.
-
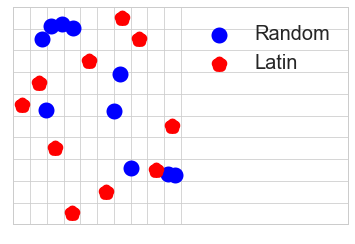
Model-free designs: These designs define rules to spread the experiments as much as possible across the input domain. Drawing points at random or in a grid are the most naive way of doing so. Other more elaborate approaches are low discrepancy sequences that try to induce some negative correlation in the selected points to spread them evenly. Some examples are Latin hyper-cube sampling and Sobol sequences.
-
Model-based designs: In these designs a feedback loop is used between some ‘optimal’ statistical criterion to collect points and a model. In general, the criterion aims to reduce some type of variance in the model. The type of optimality refers to which type of uncertainty that is reduced. For instance, a D-optimal design aims to maximize the differential Shannon information content of the model parameter estimates; an I-optimal design seeks to minimize the average prediction variance over the entire design space. See [1] for a general review on experimental design of these type with Bayesian modes.
Gaussian processes have a long tradition of being the ‘model of choice’ for designing experiments [2]. Next, we explain how you can use them in Emukit for this purpose. Of course you can generalize these ideas to other models too.
We start by loading the Branin function. We define the input space to be [-5,10]x[0,15].
from emukit.test_functions import branin_function
from emukit.core import ParameterSpace, ContinuousParameter
f, _ = branin_function()
parameter_space = ParameterSpace([ContinuousParameter('x1', -5, 10),
ContinuousParameter('x2', 0, 15)])
Emukit supports both model-free and model-based experimental design strategies. To start, as we don’t have any other information about the function, we first collect 20 points using a Latin design.
from emukit.core.initial_designs.latin_design import LatinDesign
design = LatinDesign(parameter_space)
num_data_points = 20
X = design.get_samples(num_data_points)
As they are often used to bootstrap the model, we call such model-free designs “initial”.
Now we evaluate the function at the selected points and we fit a model with GPy.
from GPy.models import GPRegression
from emukit.model_wrappers import GPyModelWrapper
Y = f(X)
model_gpy = GPRegression(X,Y)
model_emukit = GPyModelWrapper(model_gpy)
We can use the model to decide which are the best points to collect using some data collection criteria (that we call acquisition). Here we use the model variance as the acquisition function. It is known that when using Gaussian processes, selecting points of maximum variance is equivalent to maximizing the mutual information between the model and the new set of points [3] so this is a simple but mathematically grounded approach.
from emukit.experimental_design.acquisitions import ModelVariance
model_variance = ModelVariance(model = model_emukit)
As we do in other parts of Emukit, we can put our Gaussian process model to work in a data collection loop. In this case we define experiments in which 5 points are collected in parallel.
from emukit.experimental_design import ExperimentalDesignLoop
expdesign_loop = ExperimentalDesignLoop(model = model_emukit,
space = parameter_space,
acquisition = model_variance,
batch_size = 5)
By passing the function to evaluate, for instance if we are running experiments with a simulator, we can totally automate the data collection process. Let’s run 10 experiments with 5 points each.
max_iterations = 10
expdesign_loop.run_loop(f, max_iterations)
If you are running physical experiments you can just run one iteration, collect the 5 points and repeat the process over for the next batch.
Check our list of notebooks and examples if you want to learn more about how to do experimental design and other methods with Emukit. You can also check the Emukit documentation.
We’re always open to contributions! Please read our contribution guidelines for more information. We are particularly interested in contributions regarding examples and tutorials.
References on experimental design
-
[1] Kathryn Chaloner and Isabella Verdinelli, (1995). Bayesian Experimental Design: A Review, Statistical Science Vol. 10, No. 3, pp. 273-304, 1995.
-
[2] Jerome Sacks, William J Welch, Toby J Mitchell and Henry P Wynn (1989). Design and analysis of computer experiments, Statistical science, Vol. 4, No. 4, pp. 409-423, 1989.
-
[3] Niranjan Srinivas, Andreas Krause, Sham Kakade, Matthias Seeger - Gaussian process optimization in the bandit setting: No regret and experimental design, Proceedings of the 27 th International Conference on Machine Learning, Haifa, Israel, 2010.